Leveraging Animal Models to Combat Drug Resistance in Cancer Therapy
Cancer, globally recognized as a significant threat to public health, can be effectively controlled in its early stages through chemotherapy, targeted therapy, and immunotherapy, all of which inhibit tumor growth. However, an inevitable challenge emerges over time as tumors develop drug resistance, resulting in treatment failure. Unraveling the underlying mechanisms of resistance is pivotal in overcoming this hurdle and in the development of successful therapeutic strategies.
1. The mechanisms of tumor resistance
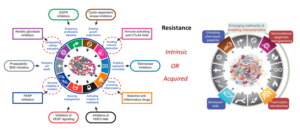
Figure 1. Relationship between cancer hallmarks and drug-resistance
In 2000, Douglas Hanahan and Robert A. Weinberg initiated the concept of cancer hallmarks, defining the six critical traits of tumor cells which encompass proliferation, invasion, and metastasis [1]. This fundamental framework evolved further in 2011 to incorporate cellular energetics—now more expansively referred to as reprogramming cellular metabolism—and the evasion of immune destruction [2]. In 2021, the spectrum broadened to propose additional hallmarks and enabling characteristics, including the unlocking of phenotypic plasticity, non-mutational epigenetic reprogramming, polymorphic microbiomes, and senescent cells [3]. This expanded perspective transcends the description of tumor cells alone, incorporating the intricacies of the tumor microenvironment (TME) (Figure 1).
The research and development of new drugs are fundamentally tied to a comprehensive understanding of tumor biology. For instance, the introduction of therapeutics such as VEGF inhibitors targeting angiogenesis and EGFR inhibitors aimed at tumor proliferation signals, have contributed significantly to advancements in tumor treatment. However, some patients may not benefit from these drug therapies due to inherent resistance. Moreover, for patients who initially respond positively, the drugs may eventually lose their efficacy over time, leading to acquired drug resistance.
Any changes in the hallmarks of cancer could potentially instigate drug resistance. This resistance is intricately tied to several factors within the tumor microenvironment, including the presence of infiltrating immune and inflammatory cells, cancer-associated fibroblasts, and vascular endothelial cells. Meanwhile, factors inherent to the tumor cells themselves, such as tumor cell heterogeneity, alterations in drug transport and metabolism, activation of alternative pathways, epigenetic changes, and the process of epithelial-mesenchymal transition, also play a significant role in the onset of drug resistance [4] (Figure 2).

Figure 2. Potential mechanisms related to drug resistance in cancer [4]
2. Strategies to establish acquired drug resistant models
Aiming to emulate real-world clinical scenarios of tumor resistance and to bolster the research and development of innovative anti-cancer drugs, WuXi Biology’s Oncology and Immunology Unit has established preclinical drug resistance models. These models leverage in vivo and in vitro methodologies (Figure 3).
2.1 In vivo Methods
The initial approach involves sourcing drug-resistant tumor tissues from patients post-surgery and directly inoculating these tissues into immunodeficient mice, establishing an in vivo model. When the same drug is administered to tumor-bearing mice, the mice exhibit the similar resistance to the therapeutic, successfully establishing a drug-resistant patient-derived xenograft (PDX) model. This drug-resistant PDX model preserves the heterogeneity and complexity typically observed in clinical tumors. However, it faces challenges such as a limited supply of clinical samples and difficulties in their acquisition. Moreover, the intricate mechanisms of drug resistance could potentially result in the loss of drug resistance in immunodeficient mice.
In the second approach, drug-sensitive tumor models including cell line derived and patient-derived xenografts are selected. These tumor-bearing mice are subjected to long-term drug treatment coupled with ongoing in vivo passaging of tumor tissues. Initially, the tumors in the treated mice exhibit slow growth. However, over time, their growth rate hastens and their sensitivity to the drugs declines, thereby indicating the successful creation of a drug-resistant tumor model (Figure 3). This method accurately simulates the process in which patients develop drug resistance following long-term treatment, as observed in clinical environments. Importantly, the resistant tumor tissues generated are not limited to the tumor cells but also include fibroblasts, immune cells, and the extracellular matrix. This model allows for a comprehensive study of the interactions between various cell types within the TME. For drugs that have not yet shown resistance in clinical scenarios or those with unclear resistance mechanisms, in vivo induction of resistance can offer proactive guidance for future novel drug development strategies. The downside of this approach is its time-intensive nature, typically necessitating 1-2 years to establish such drug-resistant models.
2.2 In vitro Methods
Approaches to developing drug-resistant models in vitro fall under two categories. The first involves the selection of sensitive tumor cell lines, where continuous drug pressure is applied within the in vitro culture system. As drug tolerance strengthens, the drug concentration progressively increases until the cells exhibit a pronounced drug-resistant phenotype. This strategy facilitates rapid in vitro screening experiments, with the mechanisms of drug resistance primarily concentrating on transformations within the tumor cells themselves. However, a limitation exists: drug-resistant cell lines induced in vitro do not ensure a 100% successful transfer in vivo. Some drug-resistant cells fail to form tumors in mice or exhibit suboptimal drug resistance.
The second in vitro method involves creating genetically engineered cell lines based on drug resistance mechanisms reported in scientific literature, such as target mutations, overexpression, and fusion proteins. These mechanisms are closely tied to clinical scenarios and can be employed for in vitro screening experiments. However, this method’s main constraint is its significant reliance on established drug resistance mechanisms and advanced genetic engineering techniques (Figure 3).
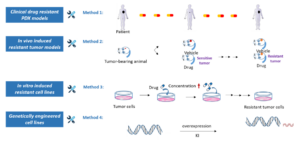
Figure 3. Strategies to establish preclinical drug-resistant tumor models
The Oncology and Immunology Unit of WuXi Biology has successfully developed over 40 drug-resistant tumor models, covering 16 popular targets such as KRAS, AR, CDK4/6, and EGFR, for supporting pre-clinical drug development (Table 1).
Table 1. Drug-resistant tumor models developed by the Oncology and Immunology Unit of WuXi Biology


3. Case study: In vitro induced T-DM1 resistant HCC1954 tumor model
Antibody-drug conjugates (ADCs), which combine monoclonal antibodies with cytotoxic drugs, are known for their high target specificity and minimal side-effects. Trastuzumab emtansine (T-DM1), the first ADC to enter the market for breast cancer treatment, is extensively used in clinical settings. However, as time progresses, patients may develop resistance to T-DM1, resulting in lower treatment response rates. To investigate the potential mechanisms behind T-DM1 resistance, we established a T-DM1-resistant HCC1954 cell line in vitro. Experimental results revealed no significant morphological differences between the parental and resistant cells. Moreover, the resistant HCC1954/T-DM1-R cell line demonstrated resistance to T-DM1, but not to its payload DM1 (Figure 4).
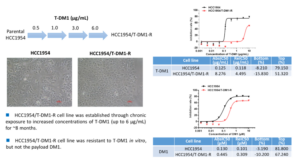
Figure 4. Establishment of in vitro induced T-DM1 resistant HCC1954 cell line
The resistant cell line was subsequently injected into mice. The resulting tumor growth was slow, demonstrating poor tumorigenicity. This led to further screening of dominant cell lines through in vivo tumorigenicity in mice. After the cells were re-isolated for in vitro validation, the xHCC1954/T-DM1-R cell line was found to be resistant. Concurrently, the resistance of xHCC1954/T-DM1-R to another ADC molecule, Trastuzumab deruxtecan (DS-8201), was evaluated under the same conditions. The results indicated that compared to the parental cell line, DS-8201 also exhibited significant resistance (Figure 5).
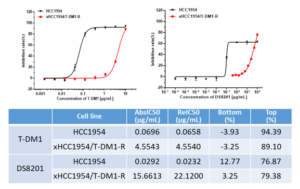
Figure 5. In vitro validation of drug resistance in xHCC1954/T-DM1-R cell line
Finally, the team performed in vivo validation experiment on the xHCC1954/T-DM1-R resistant cell line. The tumor growth curve revealed that, compared to the parental cells, the resistant cells demonstrated resistance to 5 mg/kg T-DM1, with no observable inhibition of tumor growth. Under the same conditions, the team evaluated the anti-tumor efficacy of DS-8201 in the resistant model. Initially, DS-8201 showed limited efficacy, but it significantly inhibited tumor growth during the later stages of administration (Figure 6).
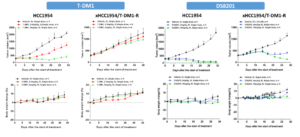

Figure 6. In vivo validation of drug resistance in xHCC1954/T-DM1-R cell line
Beyond the T-DM1 resistant cell line, the team has recently established tumor models resistant to DS-8201, utilizing in vitro induction and genetic engineering methods.
References
- Douglas Hanahan, Robert A. Weinberg. The Hallmarks of Cancer. Cell. 2000; 100(1): 57-70.
- Douglas Hanahan, Robert A. Weinberg. Hallmarks of Cancer: The Next Generation. Cell. 2011; 144(5): 646-674.
- Douglas Hanahan. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022; 12: 31-46.
- Wang X, Zhang H, Chen X. Drug resistance and combating drug resistance in cancer. Cancer Drug Resist. 2019;2(2):141-160.
- Awad, Mark M et al. “Acquired Resistance to KRASG12C Inhibition in Cancer.” The New England journal of medicine vol. 384,25 (2021): 2382-2393.
WuXi AppTec | Tumor Models:
Related Content
Unlike conventional protein-targeted therapies, which often provide only transient effects, nucleic acid therapeutics have the potential for long-lasting effects and...
VIEW RESOURCETargeted protein degradation (TPD) approaches, including molecular glue degraders (MGDs) and proteolysis targeting chimeras (PROTACs), can overcome traditional occupancy-based inhibitor...
VIEW RESOURCE
