Establishment and Optimization of scRNA-seq assay to find the mechanism of immune therapy against tumors
Tumors are complex ecosystems composed of different cell types with different phenotypes, status and gene profiles. Commonly used GEP tools like bulk RNA Sequencing can only display gene expression profiles as a whole, and cannot reflect the heterogeneous tumor cell change or immune composition in a tumor. Single cell RNA sequencing is a good tool to implement for the single cell level. However, the method and experimental system should be optimized to make sure the result is reliable and interpretable. Thus, we tried to develop and optimize our scRNA-seq assay through a series of validation steps to ensure the reliability of results for finding the mechanisms of immune therapy against tumors.
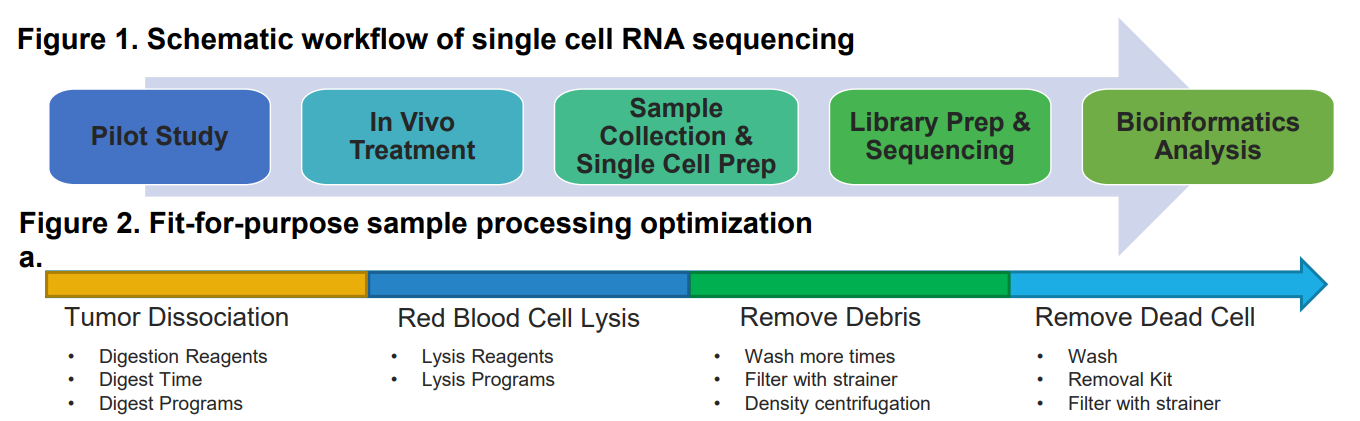
In this study, we established a reliable system for scRNA-seq analysis to elucidate the mechanism of anti-PD-1 treatment, and the interaction between tumor and immune cells. We optimized the system from the following three aspects: 1) sample processing method to confirm the cell yield, viability, and no bias introduction, 2) Library size sensitive enough for our sub-population analysis, and 3) determining whether to sort immune cells before library construction. Using this system, we tried to elucidate the mechanism of anti-PD-1 against tumors by comparing the change in scRNA-seq profiles after the treatment against two breast cancer models, EMT-6 and 4T1, which are responsive and non-responsive to anti-PD-1 treatment, respectively. Both models were treated with anti-PD-1 or isotype control and tumors were collected for scRNA-seq analysis. Interestingly, changes in both tumor cells and tumor micro-environment were observed, and this will help us to better understand how this immune therapy is able to shake up the complex ecosystem.

2021 AACR_scRNA-seq_2721_Qiyao
Related Content
WuXi AppTec provides integrated and cost-effective services for: Polyclonal and monoclonal antibodies Anti-drug antibodies (ADA), host cell protein (HCP) antibodies,...
VIEW RESOURCEWuXi AppTec scientists contributed to a research article in the journal Translational Medicine Communications which characterized the antitumor immune responses...
VIEW RESOURCE
